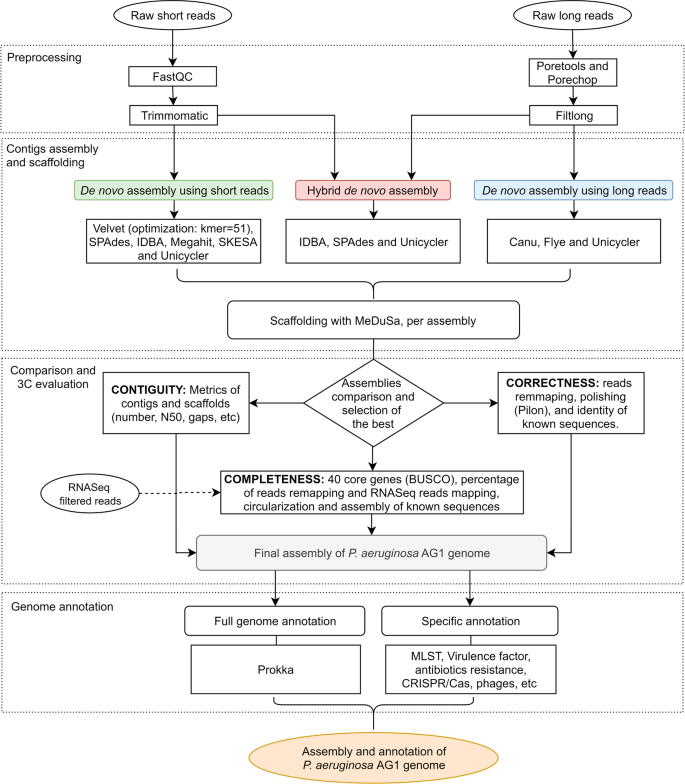
High quality 3C de novo assembly and annotation of a multidrug resistant ST-111 Pseudomonas aeruginosa genome: Benchmark of hybrid and non-hybrid assemblers | Scientific Reports
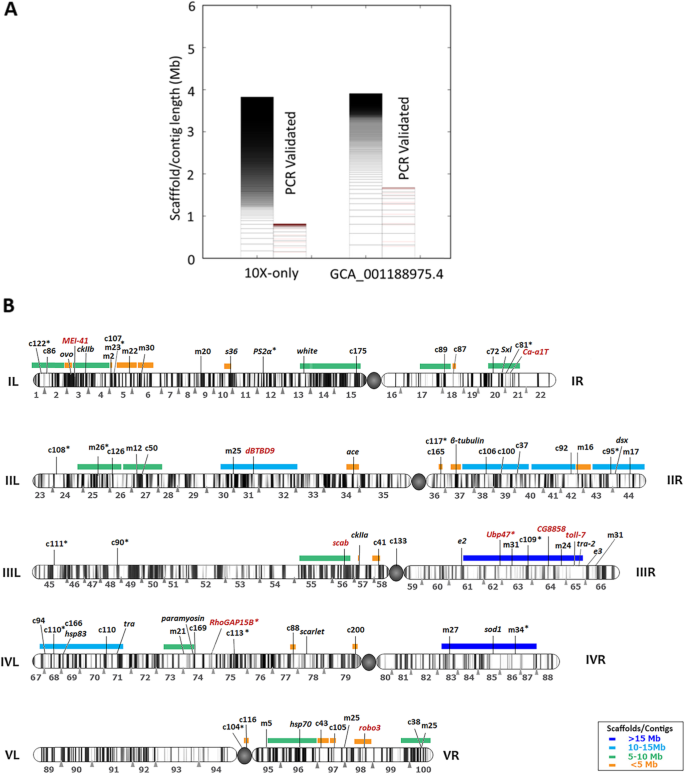
De novo assembly of the olive fruit fly ( Bactrocera oleae ) genome with linked-reads and long-read technologies minimizes gaps and provides exceptional Y chromosome assembly | SpringerLink
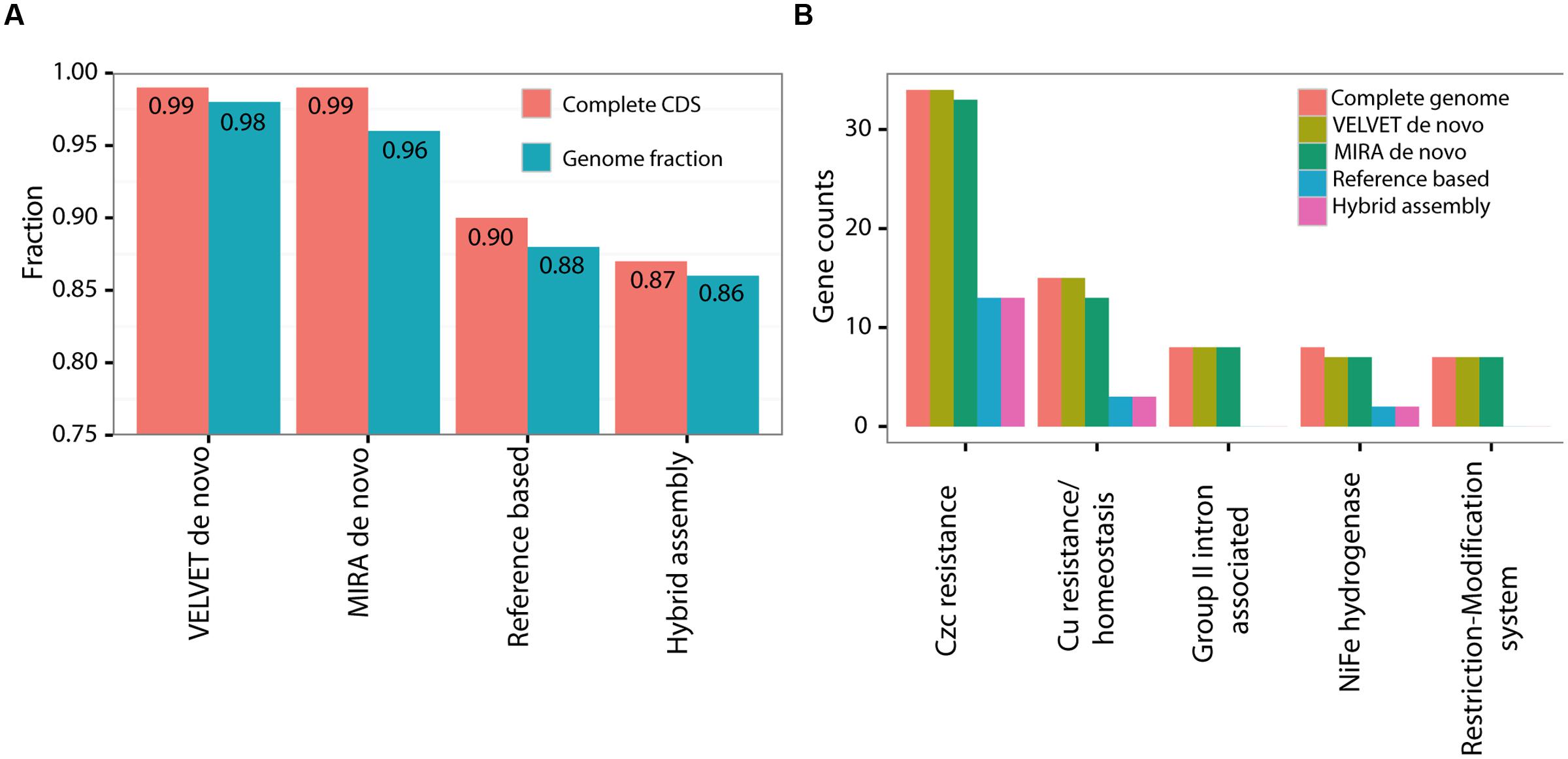
Frontiers | Why Close a Bacterial Genome? The Plasmid of Alteromonas Macleodii HOT1A3 is a Vector for Inter-Specific Transfer of a Flexible Genomic Island | Microbiology
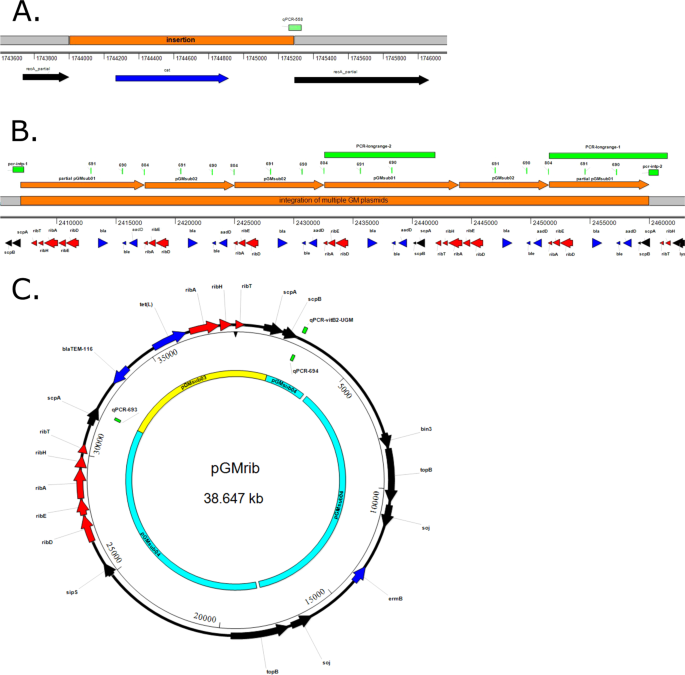
Combining short and long read sequencing to characterize antimicrobial resistance genes on plasmids applied to an unauthorized genetically modified Bacillus | Scientific Reports

A post-assembly genome-improvement toolkit (PAGIT) to obtain annotated genomes from contigs. - Abstract - Europe PMC
Semi-Automatic In Silico Gap Closure Enabled De Novo Assembly of Two Dehalobacter Genomes from Metagenomic Data

PlasFlow: predicting plasmid sequences in metagenomic data using genome signatures. - Abstract - Europe PMC
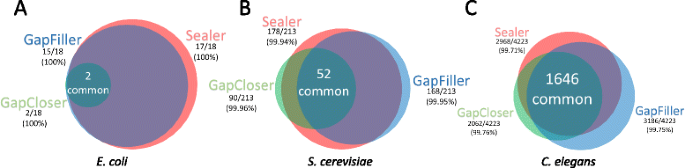
Sealer: a scalable gap-closing application for finishing draft genomes | BMC Bioinformatics | Full Text
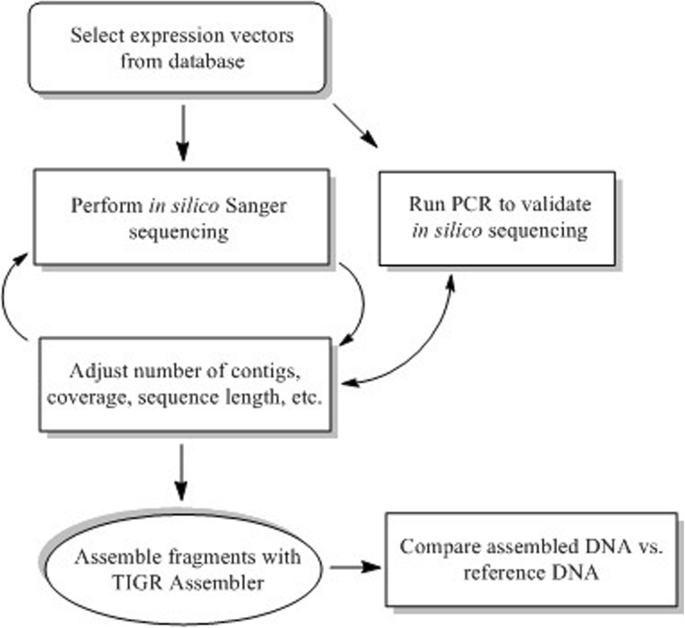
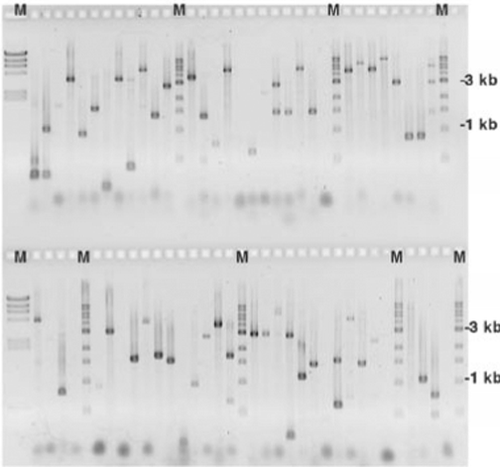
Modeling of shotgun sequencing of DNA plasmids using experimental and theoretical approaches | BMC Bioinformatics | Full Text